Inside the Tylenol Pathway: An Interactive Lesson in Why Biology Is Hard
Acetaminophen — the drug you know as Tylenol — is one of the most thoroughly mapped molecules in human biology. Every atom of its journey through the liver has been published, modeled, and picked apart for decades. It is also, paradoxically, one of the leading causes of acute liver failure in the United States.
That tension makes it the perfect teaching example for why pathway visibility matters in drug discovery.
The three pathways
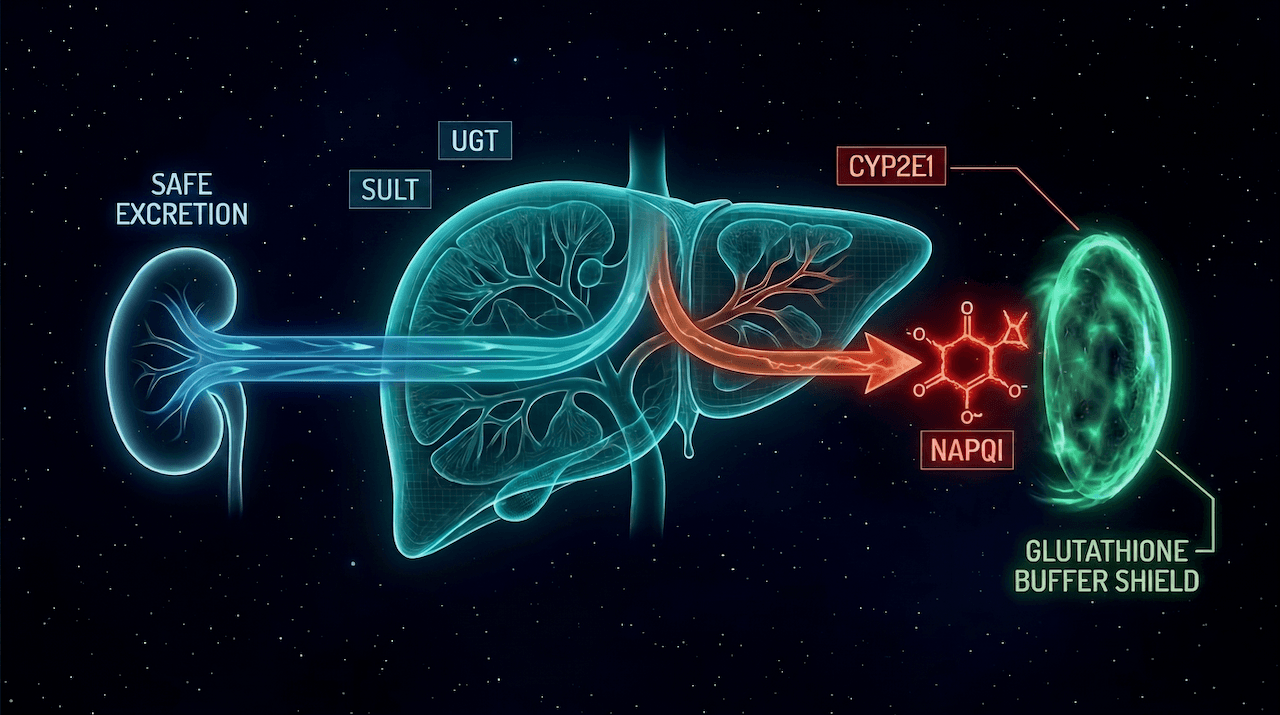
When you swallow a Tylenol, your liver routes the molecule through three parallel pathways. Two are safe. One is a ticking clock.
- Glucuronidation (~50–60%). An enzyme called UGT attaches a bulky sugar group (glucuronic acid) to acetaminophen. The result is water-soluble and harmless. Your kidneys flush it out.
- Sulfation (~30%). An enzyme called SULT attaches a sulfate group instead. Same outcome — soluble, non-toxic, excreted.
- CYP450 oxidation (~5–10%). A smaller fraction gets routed to CYP2E1, which oxidizes the molecule. The product is NAPQI — short for N-acetyl-p-benzoquinone imine — and it's toxic.
Under normal conditions, the NAPQI gets neutralized almost instantly by glutathione (GSH), the liver's antioxidant buffer, and you never notice it existed.
What breaks at overdose
Here's the part nobody tells you when they hand you the bottle.
The safe pathways — glucuronidation and sulfation — have a ceiling. They're saturable. Above about 4 grams in a single dose, they max out. They cannot process any more, no matter how badly your body wants them to.
Every molecule above that ceiling is forced down the CYP450 pathway, which means every extra milligram becomes NAPQI.
Then the glutathione buffer gets consumed. One NAPQI molecule to one glutathione molecule. When GSH runs out, NAPQI starts attacking your liver cells directly. That's acute hepatotoxicity. That's why people die.
See it in motion
Here's the system. Move the dose slider up. Watch the glucuronidation and sulfation particles saturate, the CYP2E1 branch widen with red NAPQI, the GSH buffer shrink, and the liver node flip from healthy to necrotic.
Then drop GSH to 20%. Then turn on NAC rescue. Then combine them.
Interactive Simulator
Acetaminophen Metabolic Pathway
Drag the sliders to see how dose and glutathione levels drive the flux through each pathway.
N-acetylcysteine replenishes glutathione and is the clinical antidote for acetaminophen overdose.
Try these scenarios
What to notice
A few things the simulator makes visible that prose can't:
- The safe pathways aren't lazy at low doses — they scale up with the dose until they can't. You see them accept more flux until they hit the ceiling, then stop.
- CYP2E1 is always active, at every dose. A small red trickle runs through the NAPQI branch even at 1 gram. It just gets neutralized invisibly.
- Glutathione depletion is a second failure mode, not an alternative one. Even at a moderate dose, if your GSH is already depleted — by fasting, alcohol, chronic illness — you can still hit necrosis. The simulator shows this when you drop GSH below 30% at 4g.
- NAC works by restoring GSH capacity. Turning it on doesn't remove NAPQI — it just lets the buffer catch up. Which is exactly why there's a time window for the clinical antidote.
Why this matters for AI drug discovery
This simulator runs on six lines of real pharmacology. It models three saturable pathways, a 1:1 neutralization reaction, and a damage threshold. That's enough to teach the whole story of why a "safe" drug becomes the leading cause of acute liver failure.
Now imagine an AI model trying to predict toxicity for a new molecule — one where the pathway map hasn't been fully characterized. Which enzymes metabolize it? What are their Vmax ceilings? What toxic intermediates are produced? What buffer systems neutralize them? How does the profile change across liver conditions, genetics, drug interactions?
That data mostly doesn't exist.
For acetaminophen we have the answer because fifty years of pharmacology and a thousand wet-lab experiments built the pathway map. For every new candidate molecule entering discovery, the pathway map is mostly blank. The AI is being asked to predict toxicity with a fraction of the context the Tylenol model already has.
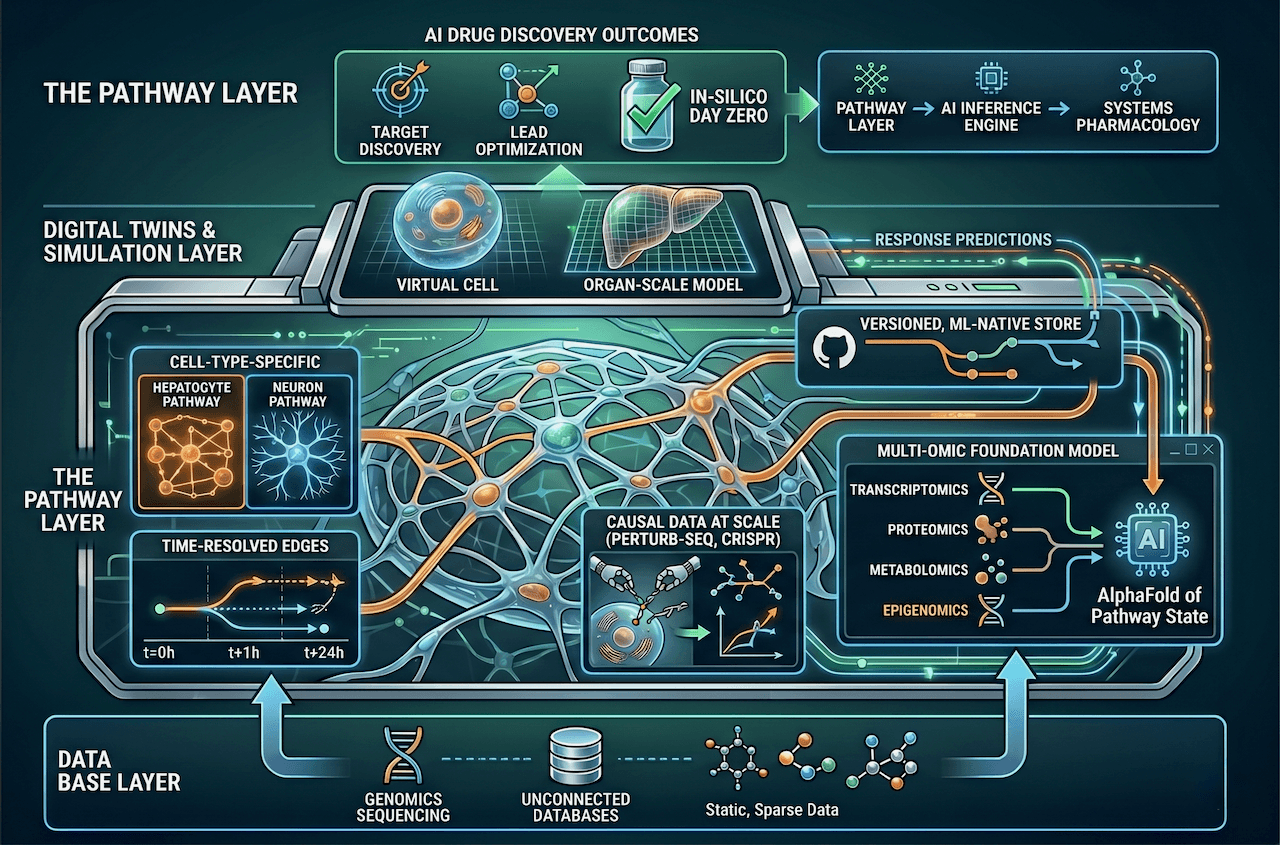
This is the point I made in The Pathway Layer: the ceiling on AI drug discovery isn't models or compute — it's the pathway data we haven't built yet. The Tylenol simulator is what pharmacology looks like when you have it. Most of biology still looks like an empty canvas.
Try it yourself
Go back up and explore the scenarios. Push the dose past 4g with full GSH and watch the system absorb it. Drop GSH to 20% and watch it fail at a therapeutic dose. Flip NAC on and watch the rescue.
Then imagine running the same simulation for ten thousand candidate molecules, each with a different pathway profile — some known, most not. That's the job AI drug discovery is trying to do.
The molecule is solved. The biology is not.
Further reading
Every reaction in this post is annotated in Reactome, the open-source curated human pathway knowledge base. Notice that you need five separate entries to assemble the full Tylenol story — which is exactly the fragmentation problem The Pathway Layer argues against.
- Glucuronidation (R-HSA-156588) — the blue pathway (~55% of a therapeutic dose, UGT1A6 is the specific isozyme for APAP)
- Cytosolic sulfonation of small molecules (R-HSA-156584) — the teal pathway (~30%, limited by PAPS availability)
- CYP2E1 monooxygenates APAP to NAPQI (R-HSA-9753285) — the canonical annotation of the toxic step
- Glutathione conjugation (R-HSA-156590) — the rescue reaction (glutathione S-transferase neutralizes NAPQI 1:1)
- Glutathione synthesis and recycling (R-HSA-174403) — the cycle that NAC therapy feeds to restore depleted GSH
Share this post