The Pathway Layer: What AI Drug Discovery Needs to Build Next
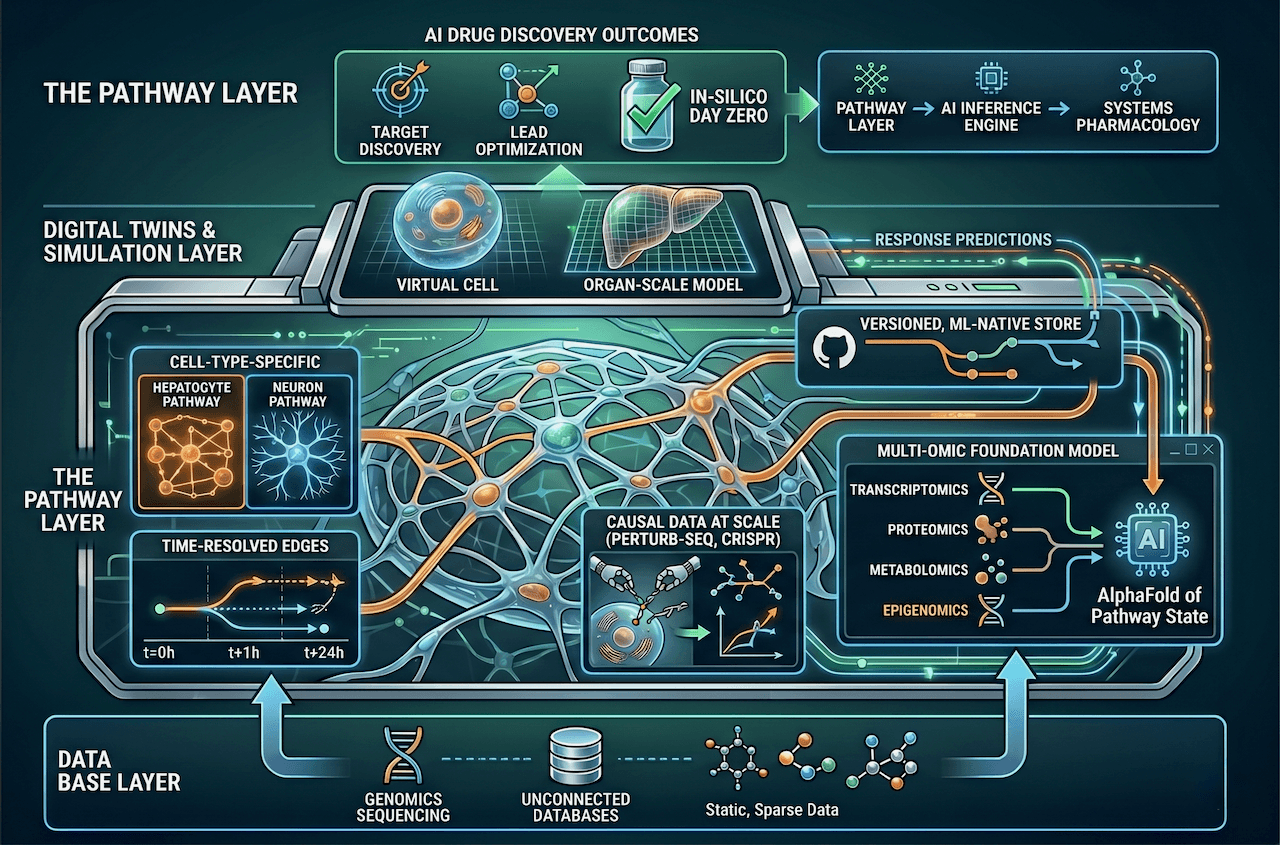
AI drug discovery has a ceiling. It isn't compute. It isn't model architecture. It isn't even data volume. The ceiling is the pathway layer — the structured, machine-readable, cell-type-specific representation of how biology actually works. And we haven't built it.
Today's AI models predict molecule properties beautifully. They predict pathway behavior poorly. The reason isn't a research gap — it's an infrastructure gap. The data those models would need either doesn't exist, doesn't connect, or sits in fifteen incompatible formats.
If we want AI drug discovery to deliver on its promise, the field has to invest in the pathway layer the way the genomics community invested in sequencing infrastructure twenty years ago.
Here's what that looks like.
1. Build pathway databases that are actually machine-readable
KEGG, Reactome, BioPAX, SBML — they exist, they're heroic, and they're not enough. Pathway maps today are:
- Cell-type agnostic — the same generic diagram for a hepatocyte and a neuron
- Static — no temporal dynamics, no signal propagation timing
- Inconsistent — the same pathway is named, segmented, and annotated differently across databases
- Sparsely curated — many pathways haven't been updated since the late 2010s
What's needed: a versioned, normalized, ML-native pathway store with cell-type stratification and time-resolved edges. Think GitHub for pathways, not PDFs.
2. Generate causal pathway data at scale, not correlation
Most pathway maps are inferred from correlations across observational data. That's a recipe for spurious edges and missing real ones. Causal data requires perturbation:
- Perturb-seq, CRISPR screens, optogenetic interventions — each one isolates a single variable
- Tens of millions of perturbations, not the tens of thousands we have today
- Across multiple cell types, not just immortalized cancer lines
The Human Cell Atlas, Tabula Sapiens, and Perturb-seq consortia are doing real work here. They need an order of magnitude more funding.
3. Multi-omics, not single-omics
A pathway's behavior at any moment depends on:
- Transcriptomics (what mRNA is being made)
- Proteomics (what protein is actually present)
- Metabolomics (what metabolites are flowing)
- Epigenomics (what's accessible to be read)
- Spatial context (where in the cell, where in the tissue)
Today's foundation models in biology mostly train on one modality. We need multi-omic foundation models — the AlphaFold of pathway state, not pathway structure.
4. Cell- and organ-level digital twins
Karr Lab's whole-cell model of Mycoplasma genitalium in 2012 was a breakthrough. Thirteen years later, we still don't have a comparable model for any human cell type. We need:
- Virtual cells that simulate a perturbation and predict downstream pathway response
- Tissue-scale models that integrate cell-cell signaling
- Organ-scale models for predicting drug effects before any wet-lab work
NVIDIA Omniverse, OpenSim, and the EU's Human Brain Project show the rendering and compute substrate exists. The biological models on top are what's missing.
5. Federated infrastructure for proprietary data
Pharma companies sit on enormous proprietary pathway data. They will never open-source it. Federated learning gives a path to train shared models without sharing raw data — exactly the architecture the field needs to break the consortium-vs-competitor deadlock.
This requires industry standards, governance frameworks, and trust infrastructure that nobody has the patience to build alone. The same pattern that makes enterprise AI work in regulated industries — auditable, permissioned, mission-aligned — applies directly here.
6. Pathway-aware foundation models
Imagine a model trained on every published perturbation experiment, every multi-omic time course, every wet-lab pathway interrogation — across cell types, species, conditions. You hand it a perturbation: "add this molecule to a hepatocyte at this concentration." It returns a predicted multi-omic state at t+1 hour, t+24 hours, t+1 week.
That model doesn't exist. It's the AlphaFold of pathway dynamics — and it's the unlock. Without it, every drug program is a blind shot at a moving target.
Who builds this
This isn't a single-company opportunity. The pathway layer is infrastructure — like sequencing was, like the internet was. It requires:
- NIH and EU funding programs specifically for pathway infrastructure, not "AI in healthcare" buzzword grants
- Open consortia (Human Cell Atlas, ENCODE, ChEMBL — but for pathways)
- Pharma open-data commitments for non-competitive precompetitive infrastructure
- A new generation of biotech companies building the picks and shovels, not the gold
The payoff
When the pathway layer exists, AI drug discovery stops being a chemistry problem dressed up in a deep learning hat. It becomes what it should have been all along: systems pharmacology with AI as the inference engine. Predictions become testable. Off-target effects become predictable. Failures shift from Phase II in humans to Day Zero in silico.
The window
The next decade of drug discovery will be decided by who builds the pathway layer first. The companies still racing to train bigger molecule models are optimizing the wrong variable.
The molecule was never the bottleneck. The biology was. It still is.
Share this post